长达几万年间,疟疾寄生虫和人类双方的基因组展开“激战”。如今,美国宾夕法尼亚大学遗传学家和由科学家组成的国际研究小组展开合作,并发现了人类基因组向“对方”发起的反击方式。
国际小组的负责人之一是来自佩雷尔曼医学院遗传研究所(Penn's Perelman School)的Sarah Tishkoff教授,他们研究的对象是15个非洲后裔族群,通过遗传分析,观察到:针对疟疾寄生虫,差异基因引起了不同程度的易感性。
相关的研究发表在6月2日美国的 Human Genetics期刊上,研究经费由Human Frontiers in Science项目提供。
疟疾仍然是全球最致命的疾病之一,每年约有一万多人死亡,其中90%的人生活在非洲。不同的人对引起疟疾的寄生虫表现不同反应;研究小组在大规模的人群中比对与疟疾侵入红细胞能力相关的一对基因。
医学院遗传所KO博士后(国际研究小组成员)称,如果试图去识别与疾病易感性相关的变异基因,重要的一步是安排适度规模的人群,不同的人群独立进化,因此,它们会出现特有的基因突变。
疟疾的生命周期需要在感染后血红细胞中进行,感染的途径是吸附在红细胞表面,这也是一些突变体——如镰刀状血细胞贫血症——为什么改变细胞形状的原因,该突变方式属于进化过程中的正向选择。
血型糖蛋白A和B的基因具有95%的相同碱基序列,由于极高的相似性,它们会发生基因重组,这就意味着只要一种基因突变,另外一种基因也会“分享”。
基因转变是帮助人类在遗传物质“军备竞赛”中胜出的关键因素。
KO博士后还说道,寄生虫基因组也存在大量的突变,与人类相比,它的突变频率比较高。在“军备竞赛”中,这是一个有力的“武器”,或许它不会胜出,但它可以提高物种的多样性。
疟疾寄生虫和人类双方基因存在“对撞”现象,对这一过程的认识有利于辅助研究人员开发出药物或疫苗,从而克服疾病。
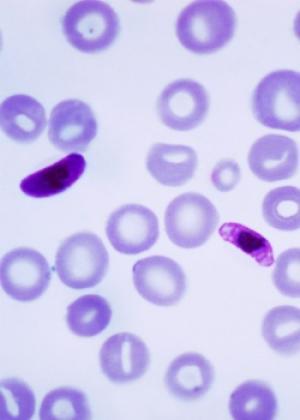
疟疾寄生虫Plasmodium falciparum
生物探索推荐英文原文
New Evidence of Genetic 'Arms Race' Against Malaria
For tens of thousands of years, the genomes of malaria parasites and humans have been at war with one another. Now, University of Pennsylvania geneticists, in collaboration with an international team of scientists, have developed a new picture of one way that the human genome has fought back.
The international team was led by Sarah Tishkoff, a Penn Integrates Knowledge professor with appointments in the genetics department in Penn's Perelman School of Medicine and the biology department of biology in the School of Arts and Sciences, and Wen-Ya Ko, a postdoctoral fellow in the genetics department at the medical school. They performed a genetic analysis of 15 ethnic groups across Africa, looking for gene variants that could explain differing local susceptibility to malaria.
Their research was published online in the journal American Journal of Human Genetics on June 2.
Malaria remains one of the deadliest diseases on the planet, annually killing about a million people, 90% of whom live in Africa. Different populations show different responses to the parasites that cause malaria; the team conducted the largest cross-population comparison ever on a pair of genes related to malaria's ability to enter red blood cells.
"When you try to identify the variants that are associated with disease susceptibility, it's important to do a very fine scale study," Ko said. "Different populations evolve independently, to a certain degree, so different populations can come up with unique mutations."
The life cycle of malaria depends on infecting red blood cells by binding to their surfaces, which is why mutations, such as sickle cell anemia, that change the overall shape of those cells are thought to have experienced positive selection.
"Both host and the parasite try to fight back with mutations; it's a co-evolution arms-race that leaves a signature of selection on the genes," Ko said. "We've identified several single-nucleotide polymorphisms that are candidates for that signature."
Across the 15 population sets, the researchers focused on polymorphisms in a pair of genes that code for proteins called glycophorin A and glycophorin B. These proteins exist on the surface of red blood cells, and changes to their shape affect the ability of the parasite causing malaria to bind to them and to infect the cells.
There are, however, two conflicting theories of why changes to glycophorin shape influence rates of malaria. One theory suggests that glycophorin A acts as a decoy, making itself more attractive to binding so that pathogens don't infect more vulnerable cells. Another theory suggests that glycophorin A mutates so that malaria parasites can't bind at all.
The researchers observed differing patterns of natural selection acting on the different regions of the two genes. They noted an excess of genetic variation being maintained in the region of glycophorin A that plays a critical role of entry of the malaria parasite into the blood cell.
"This signature of selection was strongest in populations that have the highest exposure to malaria," Tishkoff said.
In addition, the researchers identified a novel protein variant at glycophorin B in several populations with high levels of malaria that may also be a target of natural selection.
Comparisons to chimpanzee and orangutan genomes showed that these mutations occurred after the human lineage split from these closely related primates. But a process known as "gene conversion," in which similar genes can acquire mutations from one another during cell division, complicates tracking the exact history of the mutation's spread.
"The genes for glycophorin A and B arose through gene duplication. They are more than 95 percent similar to each other on the sequence level," Ko said. "Because they are so similar, sequences of A might bind to B during recombination, which means a mutation that occurs on one can be shared with the other."
That aspect of gene conversion may be a key to helping humans in the genetic arms race against malaria.
"The parasite's genome is very highly mutable, and its generation time is short, as compared to humans, so having more mutations more quickly is helpful in keeping up," Ko said. "This is one tool in the arms race. It may not win the war, but it's another way to increase variation."
A better understanding of the interplay between the genes of the malaria parasite and that of its human hosts could also give researchers an artificial advantage -- drugs or vaccines -- in the fight against the disease.
"Any new information about how malaria infects cells and how humans have evolved natural defense mechanisms against that infection adds to the body of knowledge about the pathology of malaria," Tishkoff said. "This information could aid in the development of more effective treatments against malaria."
In addition to Ko and Tishkoff, the research was conducted by Kristin A. Kaercher, Alessia Ranciaro and Jibril B. Hirbo of Penn; Emanuela Giombini and Paolo Marcatili of the Department of Biochemical Sciences at the University of Rome; Alain Froment of the Musee de l' Homme, Paris: Muntaser Ibrahim of the Department of Molecular Biology and the Institute of Endemic Diseases at the University of Khartoum; Godfrey Lema and Thomas B. Nyambo of the Department of Biochemistry at the Muhimbili University of Health and Allied Sciences in Dar es Salaam, Tanzania; Sabah A. Omar of the Kenya Medical Research Institute; and Charles Wambebe of International Biomedical Research in Africa in Abuja, Nigeria.
Their research was supported by the Human Frontiers in Science Program, the National Science Foundation and the National Institutes of Health.







